Molecular choreography of fusion proteins
We trace the refolding pathways of viral fusion proteins and pinpoint intermediate states that determine tropism and entry efficiency.
We dissect how enveloped viruses engage host membranes, refold their fusion machinery, and assemble new particles. Cryo-EM, tomography, and kinetic assays let us link structural snapshots to functional outcomes.
We trace the refolding pathways of viral fusion proteins and pinpoint intermediate states that determine tropism and entry efficiency.
Integrative EM and crosslinking strategies reveal how viral machines recruit chaperones, co-factors, and immune modulators during infection.
Mix-and-plunge workflows capture millisecond transitions so we can translate structural kinetics into mechanistic models.
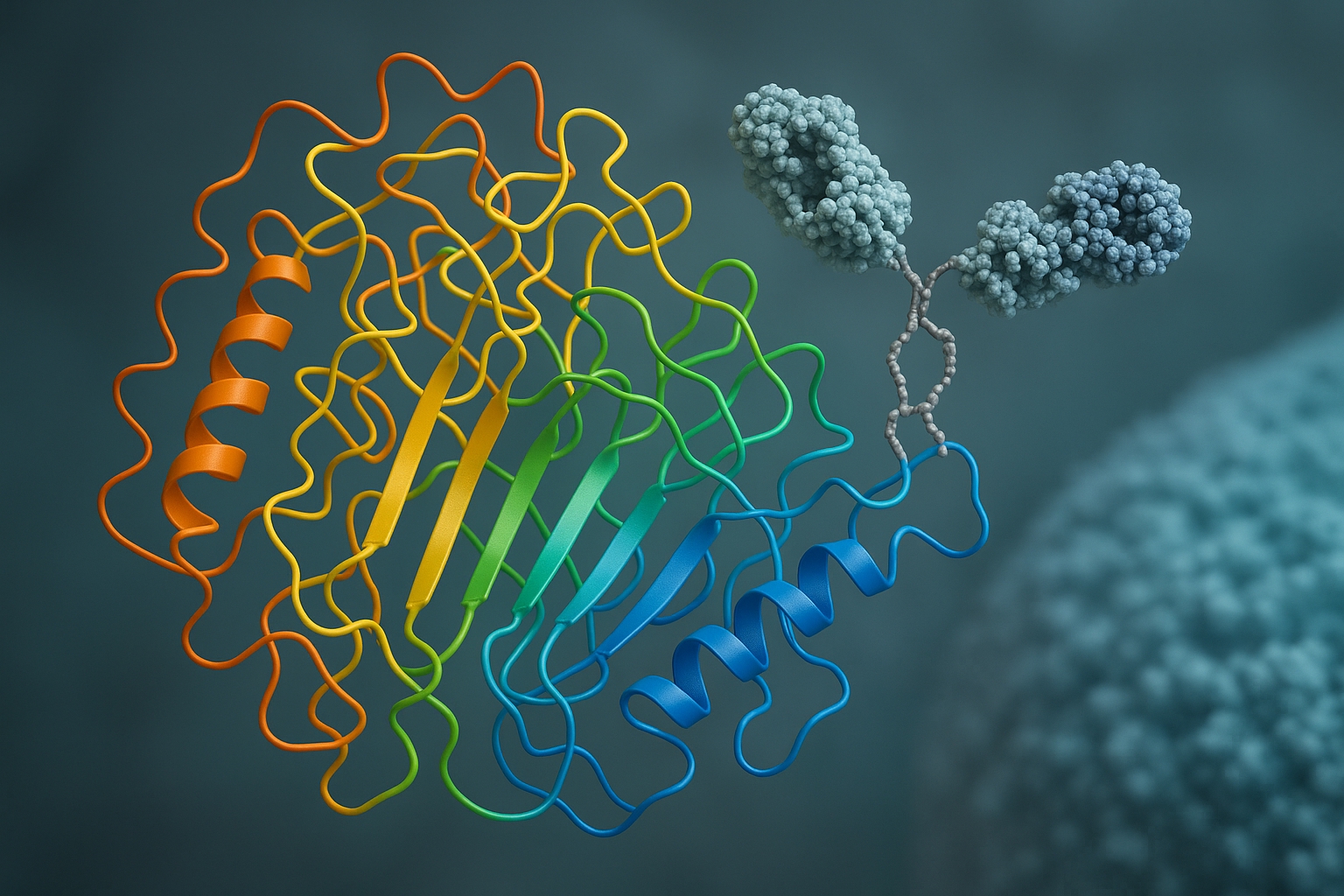
Cryo-ET combined with subtomogram averaging lets us visualize Virus X as it docks, primes, and fuses with host membranes, revealing a newly exposed pocket susceptible to inhibitory peptides.

Single-particle cryo-EM and fragment screening identify allosteric chemotypes that stall Virus Y polymerase formation and block genome replication in primary cells.